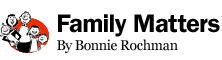Adam Foye, 11, has a mysterious disease that researchers are competing to identify using genome sequencing
For more than a decade, doctors have been trying to figure out what’s behind the weakness and fatigue that caused 11-year-old Adam Foye to miss 60 days of school last year. His symptoms match up with centronuclear myopathy (CNM), a rare muscle disease, but genetic testing shows no signs of abnormalities in any gene linked to CNM.
The puzzle may finally be solved this week. Adam, a sixth-grader in Pine Brook, N.J., is one of three patients who are the focus of an international contest to sequence their genomes and those of their parents in an effort to arrive at a diagnosis. Genome sequencing scrutinizes a person’s entire genetic code, but the contest will also analyze a subset known as the exome, looking specifically at the portion of DNA that codes for proteins. Together, both tests can offer a comprehensive look at an individual’s DNA blueprint. (Read TIME’s five-part series on the promise — and pitfalls — of sequencing children’s genomes.)
The first human genome was sequenced nearly a decade ago, but the test is still primarily used for research. It’s also beginning to be offered to patients with cancer or other diseases who are eager to know what drugs they may best respond to, as well as kids whose diagnosis remains a mystery. Traditionally, doctors have tried to figure out the cause of their illness by sequencing genes one by one or in small clusters.
But for patients like Adam, the gene-by-gene approach is too time-consuming, and his parents are hoping genome sequencing will generate answers more quickly. “This is the tedium of the current method of testing,” says Sarah Foye, Adam’s mom. “It has taken a very long time to get no answers.”
(MORE: Will My Son Develop Cancer? The Promise (and Pitfalls) of Sequencing Children’s Genomes)
As the price of sequencing drops — it’s fallen from the billions to under $10,000 — many experts believe the test will become part of routine medical care. But before that happens, there are plenty of questions that still need to be addressed. How should data be processed when results are so voluminous that they can’t even be sent electronically? How can all those results be compressed into a summary that makes sense to patients — and their doctors? And what should be done with unexpected findings — that a child has a gene, for example, that significantly increases her risk of breast cancer?
Researchers at Boston Children’s Hospital are hoping the contest they’re sponsoring will offer some guidance. It’s called Children’s Leadership Award for the Reliable Interpretation and Transmission of Your Genomic Information, or CLARITY, which is a tidy acronym for a lofty ambition. Not only do the contest’s organizers hope to find the genetic cause for each child’s disease, but the winner will provide a kind of “best practices” blueprint for reliably interpreting and sharing genomic data.
Guidelines need to be established before sequencing could go mainstream, so that doctors as well as patients understand what individual genetic changes mean. “We discovered there were no standards,” says Dr. Isaac Kohane, a CLARITY co-organizer and professor of pediatrics and health sciences and technology at Harvard Medical School. “We decided a competition would be helpful so we could compare what a clinical report would look like by multiple teams competing to sequence genomes. It takes a while for this to be reduced to reliable genomic practice.”
Kohane hopes the $25,000 prize — meager by scientific standards but still something — and the chance to help shape a powerful new technology will nudge researchers to participate. The ultimate goal: to translate all this complexity into an accurate, easily understandable script that a doctor not trained in genetics can use to make sense of the genetic jumble for an anxious mom or dad.
Sarah Foye is one of those anxious moms. Her mind is constantly churning with what-ifs: are there medical conditions associated with Adam’s disease that she’s unaware of because she doesn’t know which mutation he has? On the flip side, might the echocardiograms he gets regularly because they’re recommended for patients with CNM actually be unnecessary? And what about family planning? If she’d known from the beginning that his mutation was random, not hereditary, she may have seriously considered having another child. Pinpointing Adam’s genetic mutation could also help him understand his risks of transmitting the gene to his children.
(MORE: Why Cheaper Genetic Testing Could Cost Us a Fortune)
Adam, an above-average student and an avid video gamer (he’s obsessed with Minecraft), gave his consent to let 23 teams of contestants from around the world analyze his data. The researchers—from countries including Slovenia, Sweden, China, India and the U.S.—have scoured the information gleaned from sequencing the Foyes and the two other families participating and compared it to a database of reference genomes, to see if the kids have known diseases. They are also relying on computational models to determine if the children have new genetic changes that haven’t yet been identified or linked to disease.
It took the teams four months to interpret the kids’ genes and those of their parents’ and assemble the findings in simply worded documents that can guide patient care. The final reports were submitted at the end of September.
Sarah Foye, an occupational therapist, is eager to read them. She wants to know what types of genetic aberrations her son has, and why he gets so tired that he can’t even walk down the grocery-store aisle. He’s a pretty happy kid but is confined to using a motorized scooter in order to keep up with his friends much of the time. He’s had a feeding tube for years and isn’t strong enough to carry his own schoolbooks.
Over the years, Foye has had no choice but to be patient as test after test revealed nothing. “You’re biting your nails,” she says, “and then it comes back negative.” Her hopes are high that the 23 teams of researchers will provide an answer. She’s traveling to San Francisco, where the winner of the CLARITY contest will be announced on Wednesday at the annual meeting of the American Society of Human Genetics. “I’m dying to know what he has,” says Foye.